
Python package for concise, transparent, and accurate predictive modeling.
All sklearn-compatible and easy to use.
For interpretability in NLP, check out our new package: imodelsX

Modern machine-learning models are increasingly complex, often making them difficult to interpret. This package provides a simple interface for fitting and using state-of-the-art interpretable models, all compatible with scikit-learn. These models can often replace black-box models (e.g. random forests) with simpler models (e.g. rule lists) while improving interpretability and computational efficiency, all without sacrificing predictive accuracy! Simply import a classifier or regressor and use the fit and predict methods, same as standard scikit-learn models.
from sklearn.model_selection import train_test_split
from imodels import get_clean_dataset, HSTreeClassifierCV # import any imodels model here
# prepare data (a sample clinical dataset)
X, y, feature_names = get_clean_dataset('csi_pecarn_pred')
X_train, X_test, y_train, y_test = train_test_split(
X, y, random_state=42)
# fit the model
model = HSTreeClassifierCV(max_leaf_nodes=4) # initialize a tree model and specify only 4 leaf nodes
model.fit(X_train, y_train, feature_names=feature_names) # fit model
preds = model.predict(X_test) # discrete predictions: shape is (n_test, 1)
preds_proba = model.predict_proba(X_test) # predicted probabilities: shape is (n_test, n_classes)
print(model) # print the model
------------------------------
Decision Tree with Hierarchical Shrinkage
Prediction is made by looking at the value in the appropriate leaf of the tree
------------------------------
|--- FocalNeuroFindings2 <= 0.50
| |--- HighriskDiving <= 0.50
| | |--- Torticollis2 <= 0.50
| | | |--- value: [0.10]
| | |--- Torticollis2 > 0.50
| | | |--- value: [0.30]
| |--- HighriskDiving > 0.50
| | |--- value: [0.68]
|--- FocalNeuroFindings2 > 0.50
| |--- value: [0.42]
Installation
Install with pip install imodels (see here for help).
Supported models
🗂️ Docs 📄 Research paper 🔗 Reference code implementation
| Model | Reference | Description |
|---|---|---|
| Rulefit rule set | 🗂️, 📄, 🔗 | Fits a sparse linear model on rules extracted from decision trees |
| Skope rule set | 🗂️, 🔗 | Extracts rules from gradient-boosted trees, deduplicates them, then linearly combines them based on their OOB precision |
| Boosted rule set | 🗂️, 📄, 🔗 | Sequentially fits a set of rules with Adaboost |
| Slipper rule set | 🗂️, 📄 | Sequentially learns a set of rules with SLIPPER |
| Bayesian rule set | 🗂️, 📄, 🔗 | Finds concise rule set with Bayesian sampling (slow) |
| Bayesian rule list | 🗂️, 📄, 🔗 | Fits compact rule list distribution with Bayesian sampling (slow) |
| Greedy rule list | 🗂️, 🔗 | Uses CART to fit a list (only a single path), rather than a tree |
| OneR rule list | 🗂️, 📄 | Fits rule list restricted to only one feature |
| Optimal rule tree | 🗂️, 📄, 🔗 | Fits succinct tree using global optimization for sparsity (GOSDT) |
| Greedy rule tree | 🗂️, 📄, 🔗 | Greedily fits tree using CART |
| C4.5 rule tree | 🗂️, 📄, 🔗 | Greedily fits tree using C4.5 |
| TAO rule tree | 🗂️, 📄 | Fits tree using alternating optimization |
| Iterative random forest |
🗂️, 📄, 🔗 | Repeatedly fit random forest, giving features with high importance a higher chance of being selected |
| Sparse integer linear model |
🗂️, 📄 | Sparse linear model with integer coefficients |
| Tree GAM | 🗂️, 📄, 🔗 | Generalized additive model fit with short boosted trees |
| Greedy treesums (FIGS) | 🗂️,ㅤ📄 | Sum of small trees with very few total rules (FIGS) |
| Hierarchical shrinkage wrapper |
🗂️, 📄 | Improve a decision tree, random forest, or gradient-boosting ensemble with ultra-fast, post-hoc regularization |
| RF+ (MDI+) | 🗂️, 📄 | Flexible random forest-based feature importance |
| Distillation wrapper |
🗂️ | Train a black-box model, then distill it into an interpretable model |
| AutoML wrapper | 🗂️ | Automatically fit and select an interpretable model |
| More models | ⌛ | (Coming soon!) Lightweight Rule Induction, MLRules, … |
Demo notebooks
Demos are contained in the notebooks folder.
Quickstart demo
Shows how to fit, predict, and visualize with different interpretable modelsAutogluon demo
Fit/select an interpretable model automatically using Autogluon AutoMLQuickstart colab demo 
Shows how to fit, predict, and visualize with different interpretable models
Clinical decision rule notebook
Shows an example of usingimodels for deriving a clinical decision rule
Posthoc analysis
We also include some demos of posthoc analysis, which occurs after fitting models: posthoc.ipynb shows different simple analyses to interpret a trained model and uncertainty.ipynb contains basic code to get uncertainty estimates for a modelWhat's the difference between the models?
The final form of the above models takes one of the following forms, which aim to be simultaneously simple to understand and highly predictive:
| Rule set | Rule list | Rule tree | Algebraic models |
|---|---|---|---|
 |
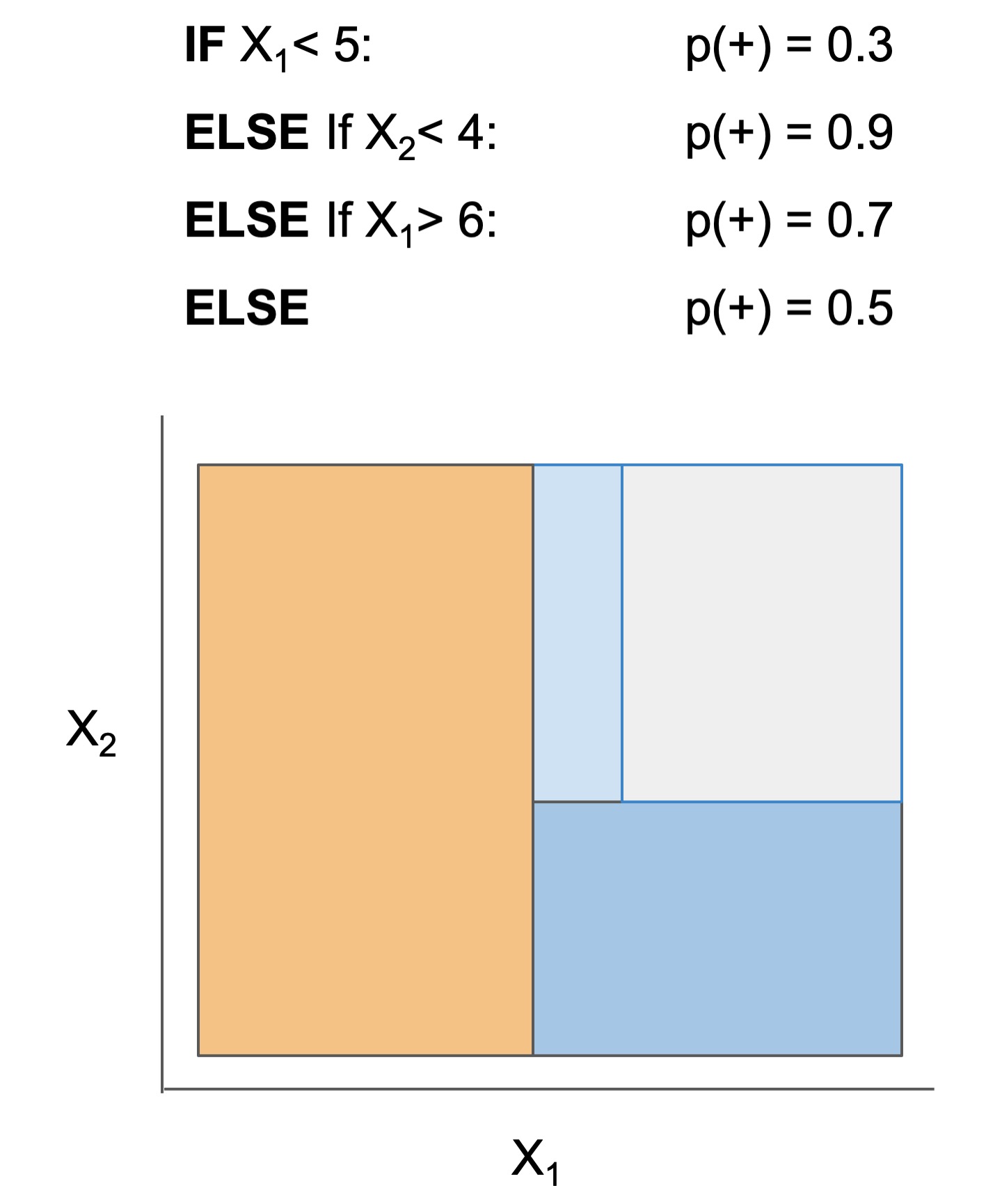 |
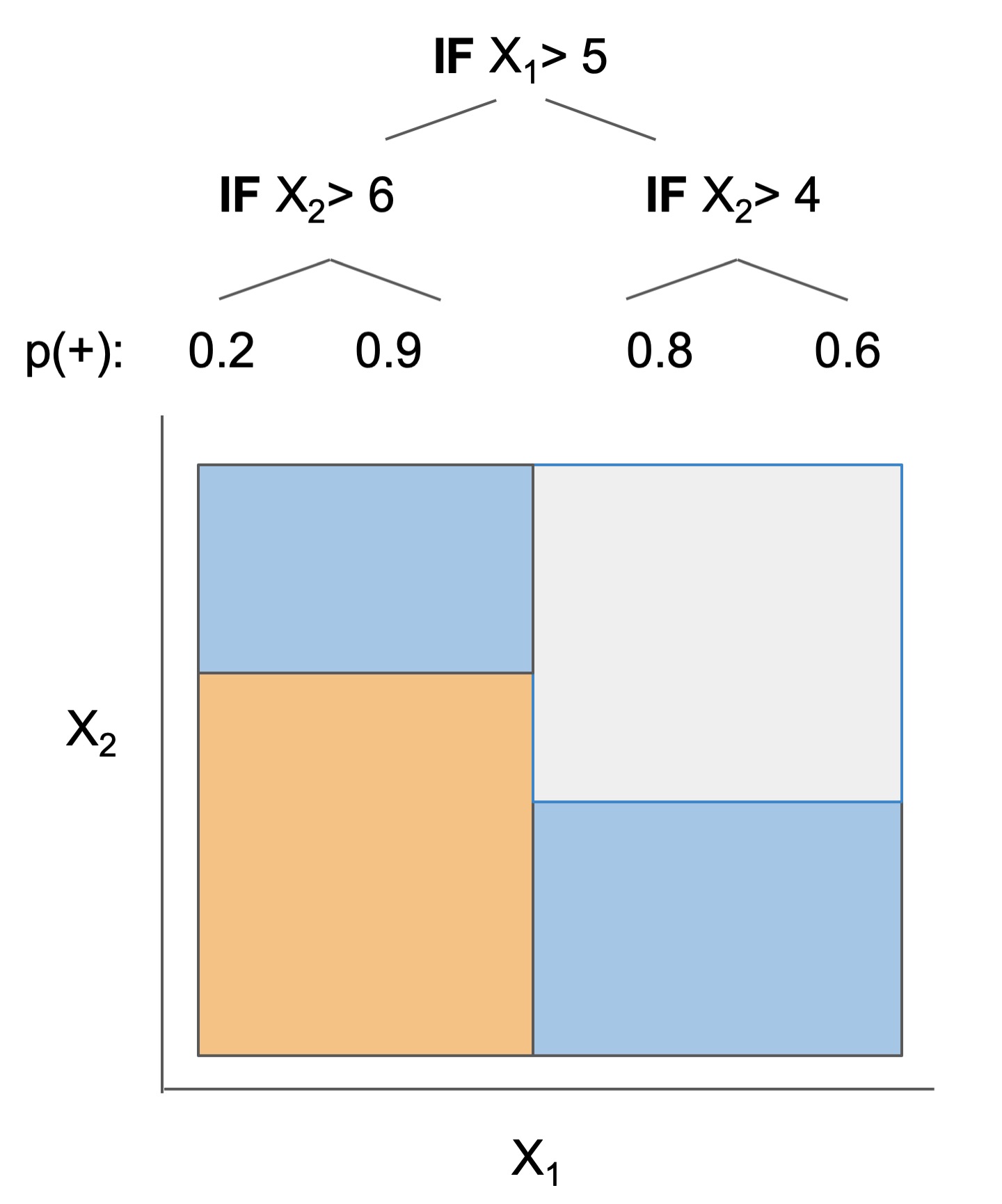 |
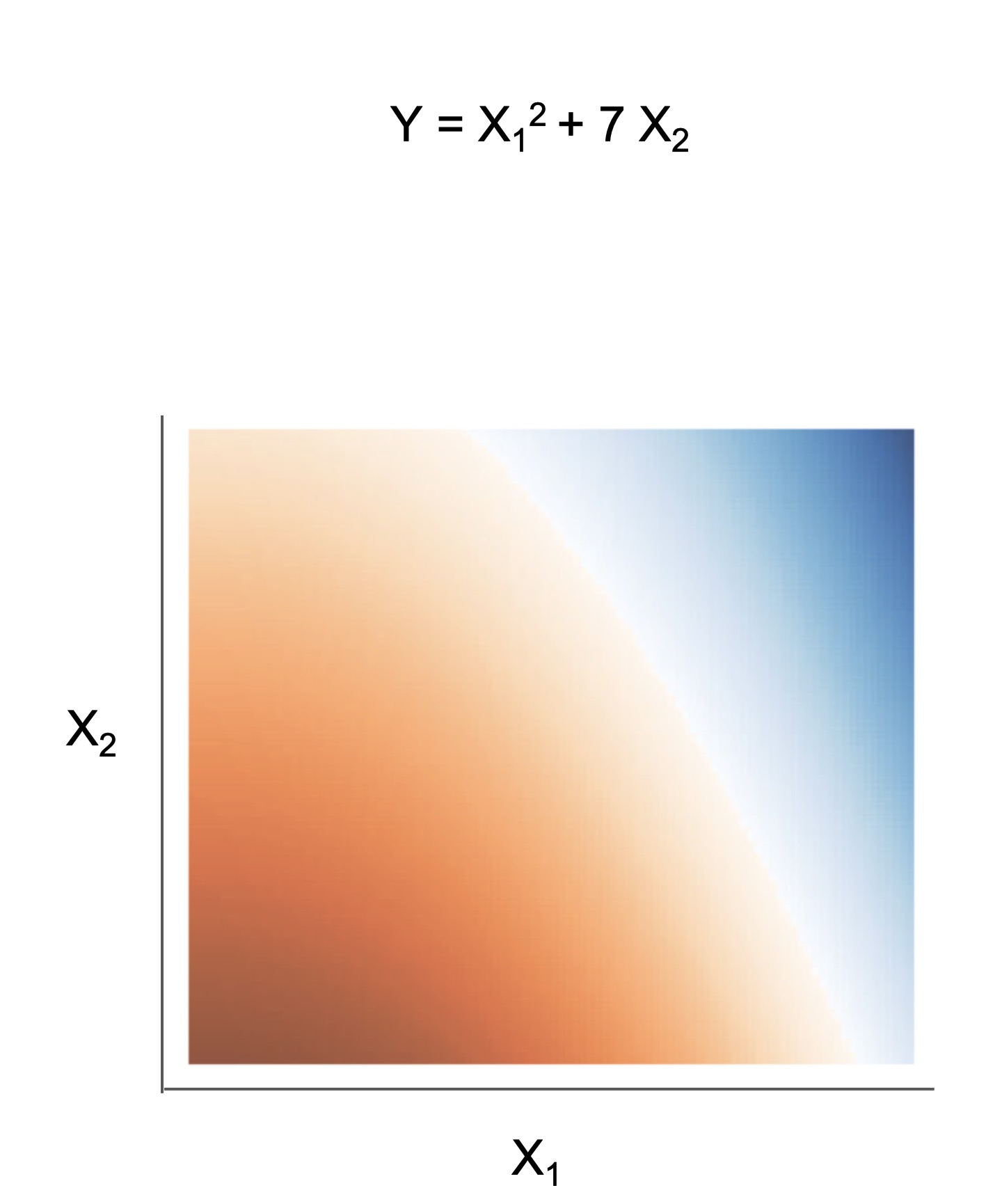 |
Different models and algorithms vary not only in their final form but also in different choices made during modeling, such as how they generate, select, and postprocess rules:
| Rule candidate generation | Rule selection | Rule postprocessing |
|---|---|---|
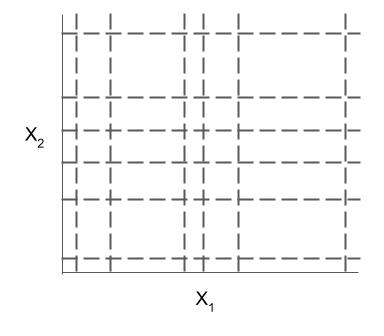 |
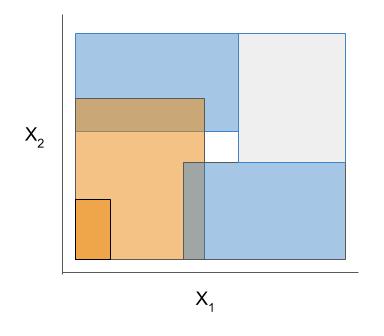 |
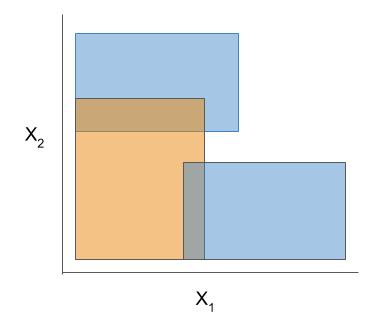 |
Ex. RuleFit vs. SkopeRules
RuleFit and SkopeRules differ only in the way they prune rules: RuleFit uses a linear model whereas SkopeRules heuristically deduplicates rules sharing overlap.Ex. Bayesian rule lists vs. greedy rule lists
Bayesian rule lists and greedy rule lists differ in how they select rules; bayesian rule lists perform a global optimization over possible rule lists while Greedy rule lists pick splits sequentially to maximize a given criterion.Ex. FPSkope vs. SkopeRules
FPSkope and SkopeRules differ only in the way they generate candidate rules: FPSkope uses FPgrowth whereas SkopeRules extracts rules from decision trees.Support for different tasks
Different models support different machine-learning tasks. Current support for different models is given below (each of these models can be imported directly from imodels (e.g. from imodels import RuleFitClassifier):
| Model | Binary classification | Regression | Notes |
|---|---|---|---|
| Rulefit rule set | RuleFitClassifier | RuleFitRegressor | |
| Skope rule set | SkopeRulesClassifier | ||
| Boosted rule set | BoostedRulesClassifier | BoostedRulesRegressor | |
| SLIPPER rule set | SlipperClassifier | ||
| Bayesian rule set | BayesianRuleSetClassifier | Fails for large problems | |
| Bayesian rule list | BayesianRuleListClassifier | ||
| Greedy rule list | GreedyRuleListClassifier | ||
| OneR rule list | OneRClassifier | ||
| Optimal rule tree (GOSDT) | OptimalTreeClassifier | Requires gosdt, fails for large problems | |
| Greedy rule tree (CART) | GreedyTreeClassifier | GreedyTreeRegressor | |
| C4.5 rule tree | C45TreeClassifier | ||
| TAO rule tree | TaoTreeClassifier | TaoTreeRegressor | |
| Iterative random forest | IRFClassifier | Requires irf | |
| Sparse integer linear model | SLIMClassifier | SLIMRegressor | Requires extra dependencies for speed |
| Tree GAM | TreeGAMClassifier | TreeGAMRegressor | |
| Greedy tree sums (FIGS) | FIGSClassifier | FIGSRegressor | |
| Hierarchical shrinkage | HSTreeClassifierCV | HSTreeRegressorCV | Wraps any sklearn tree-based model |
| Distillation | DistilledRegressor | Wraps any sklearn-compatible models | |
| AutoML model | AutoInterpretableClassifier️ | AutoInterpretableRegressor️ |
Extras
Data-wrangling functions for working with popular tabular datasets (e.g. compas).
These functions, in conjunction with imodels-data and imodels-experiments, make it simple to download data and run experiments on new models.Explain classification errors with a simple posthoc function.
Fit an interpretable model to explain a previous model's errors (ex. in this notebook📓).Fast and effective discretizers for data preprocessing.
| Discretizer | Reference | Description |
|---|---|---|
| MDLP | 🗂️, 🔗, 📄 | Discretize using entropy minimization heuristic |
| Simple | 🗂️, 🔗 | Simple KBins discretization |
| Random Forest | 🗂️ | Discretize into bins based on random forest split popularity |
Rule-based utils for customizing models
The code here contains many useful and customizable functions for rule-based learning in the util folder. This includes functions / classes for rule deduplication, rule screening, and converting between trees, rulesets, and neural networks.Our favorite models
After developing and playing with imodels, we developed a few new models to overcome limitations of existing interpretable models.
FIGS: Fast interpretable greedy-tree sums
Fast Interpretable Greedy-Tree Sums (FIGS) is an algorithm for fitting concise rule-based models. Specifically, FIGS generalizes CART to simultaneously grow a flexible number of trees in a summation. The total number of splits across all the trees can be restricted by a pre-specified threshold, keeping the model interpretable. Experiments across a wide array of real-world datasets show that FIGS achieves state-of-the-art prediction performance when restricted to just a few splits (e.g. less than 20).
Example FIGS model. FIGS learns a sum of trees with a flexible number of trees; to make its prediction, it sums the result from each tree.
Hierarchical shrinkage: post-hoc regularization for tree-based methods
📄 Paper (ICML 2022), 🔗 Post, 📌 Citation
Hierarchical shrinkage is an extremely fast post-hoc regularization method which works on any decision tree (or tree-based ensemble, such as Random Forest). It does not modify the tree structure, and instead regularizes the tree by shrinking the prediction over each node towards the sample means of its ancestors (using a single regularization parameter). Experiments over a wide variety of datasets show that hierarchical shrinkage substantially increases the predictive performance of individual decision trees and decision-tree ensembles.
HS Example. HS applies post-hoc regularization to any decision tree by shrinking each node towards its parent.
MDI+: Flexible Tree-Based Feature Importance
MDI+ is a novel feature importance framework, which generalizes the popular mean decrease in impurity (MDI) importance score for random forests. At its core, MDI+ expands upon a recently discovered connection between linear regression and decision trees. In doing so, MDI+ enables practitioners to (1) tailor the feature importance computation to the data/problem structure and (2) incorporate additional features or knowledge to mitigate known biases of decision trees. In both real data case studies and extensive real-data-inspired simulations, MDI+ outperforms commonly used feature importance measures (e.g., MDI, permutation-based scores, and TreeSHAP) by substantional margins.
References
Readings
Reference implementations (also linked above)
The code here heavily derives from the wonderful work of previous projects. We seek to to extract out, unify, and maintain key parts of these projects.- sklearn-expertsys - by @tmadl and @kenben based on original code by Ben Letham
- rulefit - by @christophM
- skope-rules - by the skope-rules team (including @ngoix, @floriangardin, @datajms, Bibi Ndiaye, Ronan Gautier)
- boa - by @wangtongada
Related packages
- gplearn: symbolic regression/classification
- pysr: fast symbolic regression
- pygam: generative additive models
- interpretml: boosting-based gam
- h20 ai: gams + glms (and more)
- optbinning: data discretization / scoring models
Updates
- For updates, star the repo, see this related repo, or follow @csinva_
- Please make sure to give authors of original methods / base implementations appropriate credit!
- Contributing: pull requests very welcome!
Please cite the package if you use it in an academic work :)
@software{
imodels2021,
title = {imodels: a python package for fitting interpretable models},
journal = {Journal of Open Source Software},
publisher = {The Open Journal},
year = {2021},
author = {Singh, Chandan and Nasseri, Keyan and Tan, Yan Shuo and Tang, Tiffany and Yu, Bin},
volume = {6},
number = {61},
pages = {3192},
doi = {10.21105/joss.03192},
url = {https://doi.org/10.21105/joss.03192},
}
Expand source code
"""
.. include:: ../readme.md
"""
# Python `imodels` package for interpretable models compatible with scikit-learn.
# Github repo available [here](https://github.com/csinva/imodels)
from .algebraic.slim import SLIMRegressor, SLIMClassifier
from .algebraic.tree_gam import TreeGAMClassifier, TreeGAMRegressor
from .algebraic.marginal_shrinkage_linear_model import (
MarginalShrinkageLinearModelRegressor,
)
from .discretization.discretizer import RFDiscretizer, BasicDiscretizer
from .discretization.mdlp import MDLPDiscretizer, BRLDiscretizer
from .experimental.bartpy import BART
from .rule_list.bayesian_rule_list.bayesian_rule_list import BayesianRuleListClassifier
# from .experimental.corels_wrapper import OptimalRuleListClassifier
from .rule_list.greedy_rule_list import GreedyRuleListClassifier
from .rule_list.one_r import OneRClassifier
from .rule_set import boosted_rules
from .rule_set.boosted_rules import *
from .rule_set.boosted_rules import BoostedRulesClassifier
from .rule_set.brs import BayesianRuleSetClassifier
from .rule_set.fplasso import FPLassoRegressor, FPLassoClassifier
from .rule_set.fpskope import FPSkopeClassifier
from .rule_set.rule_fit import RuleFitRegressor, RuleFitClassifier
from .rule_set.skope_rules import SkopeRulesClassifier
from .rule_set.slipper import SlipperClassifier
from .tree.c45_tree.c45_tree import C45TreeClassifier
from .tree.cart_ccp import (
DecisionTreeCCPClassifier,
DecisionTreeCCPRegressor,
HSDecisionTreeCCPClassifierCV,
HSDecisionTreeCCPRegressorCV,
)
# from .tree.iterative_random_forest.iterative_random_forest import IRFClassifier
# from .tree.optimal_classification_tree import OptimalTreeModel
from .tree.cart_wrapper import GreedyTreeClassifier, GreedyTreeRegressor
from .tree.figs import FIGSRegressor, FIGSClassifier, FIGSRegressorCV, FIGSClassifierCV
from .tree.gosdt.pygosdt import OptimalTreeClassifier
from .tree.gosdt.pygosdt_shrinkage import (
HSOptimalTreeClassifier,
HSOptimalTreeClassifierCV,
)
from .tree.hierarchical_shrinkage import (
HSTreeRegressor,
HSTreeClassifier,
HSTreeRegressorCV,
HSTreeClassifierCV,
)
from .tree.tao import TaoTreeClassifier, TaoTreeRegressor
from .util.automl import AutoInterpretableClassifier, AutoInterpretableRegressor
from .util.data_util import get_clean_dataset
from .util.distillation import DistilledRegressor
from .util.explain_errors import explain_classification_errors
from .clustering.stableclustering import StableClustering
CLASSIFIERS = [
BayesianRuleListClassifier,
GreedyRuleListClassifier,
SkopeRulesClassifier,
BoostedRulesClassifier,
SLIMClassifier,
SlipperClassifier,
BayesianRuleSetClassifier,
C45TreeClassifier,
OptimalTreeClassifier,
OneRClassifier,
SlipperClassifier,
RuleFitClassifier,
TaoTreeClassifier,
TreeGAMClassifier,
FIGSClassifier,
HSTreeClassifier,
HSTreeClassifierCV,
GreedyTreeClassifier,
AutoInterpretableClassifier,
] # , IRFClassifier
REGRESSORS = [
RuleFitRegressor,
SLIMRegressor,
GreedyTreeRegressor,
FIGSRegressor,
TaoTreeRegressor,
TreeGAMRegressor,
HSTreeRegressor,
HSTreeRegressorCV,
BART,
AutoInterpretableRegressor,
]
ESTIMATORS = CLASSIFIERS + REGRESSORS
DISCRETIZERS = [RFDiscretizer, BasicDiscretizer,
MDLPDiscretizer, BRLDiscretizer]Sub-modules
imodels.algebraic-
Generic class for models that take the form of algebraic equations (e.g. linear models).
imodels.clusteringimodels.discretizationimodels.experimentalimodels.importance-
Feature importance methods for black box models
imodels.rule_list-
Generic class for models that take the form of a list of rules.
imodels.rule_set-
Generic class for models that take the form of a set of (potentially overlapping) rules.
imodels.tree-
Generic class for models that take the form of a tree of rules.
imodels.util-
Shared utilities for implementing different interpretable models.




